Synthetic Biology PhD Training Program
SynBAS is a NSF-funded National Research Traineeship (NRT) program, focused on convergent synthetic biology training for PhD graduate students. Students will learn the principles of living systems across scales–from molecules, to cells, to organisms, to communities.
SynBAS
Synthesizing Biology Across Scales: The Center’s PhD Training Program
Have you ever wondered how biology becomes technology? Even the seemingly simple example of biochemical production from sustainable feedstocks requires synthesis of diverse phenomena, including the underlying chemistry, physics, biology, engineering, math, and business, that makes technology possible. Synthetic biologists of the future will need to learn fundamentals, integrate across disciplines, and navigate challenges across scales in order to be successful.
SynBAS brings together trainees from diverse backgrounds, organizing the principles of synthetic biology along biological scales, to promote research breakthroughs at the interfaces.
SynBAS is a National Research Traineeship (NRT) program, focused on convergent synthetic biology training for PhD graduate students. Through the NRT program components, students will learn the principles of living systems across scales–from molecules, to cells, to organisms, to communities. The NRT program offers direct financial support to PhD graduate student trainees, as well as communications training, career mentoring, networking connections to academia and industry, and more. Many of the NRT’s programs and initiatives including the courses, workshops, retreats and community events, are open to all CSB students and postdocs at Northwestern.
See CSB-Wide Opportunities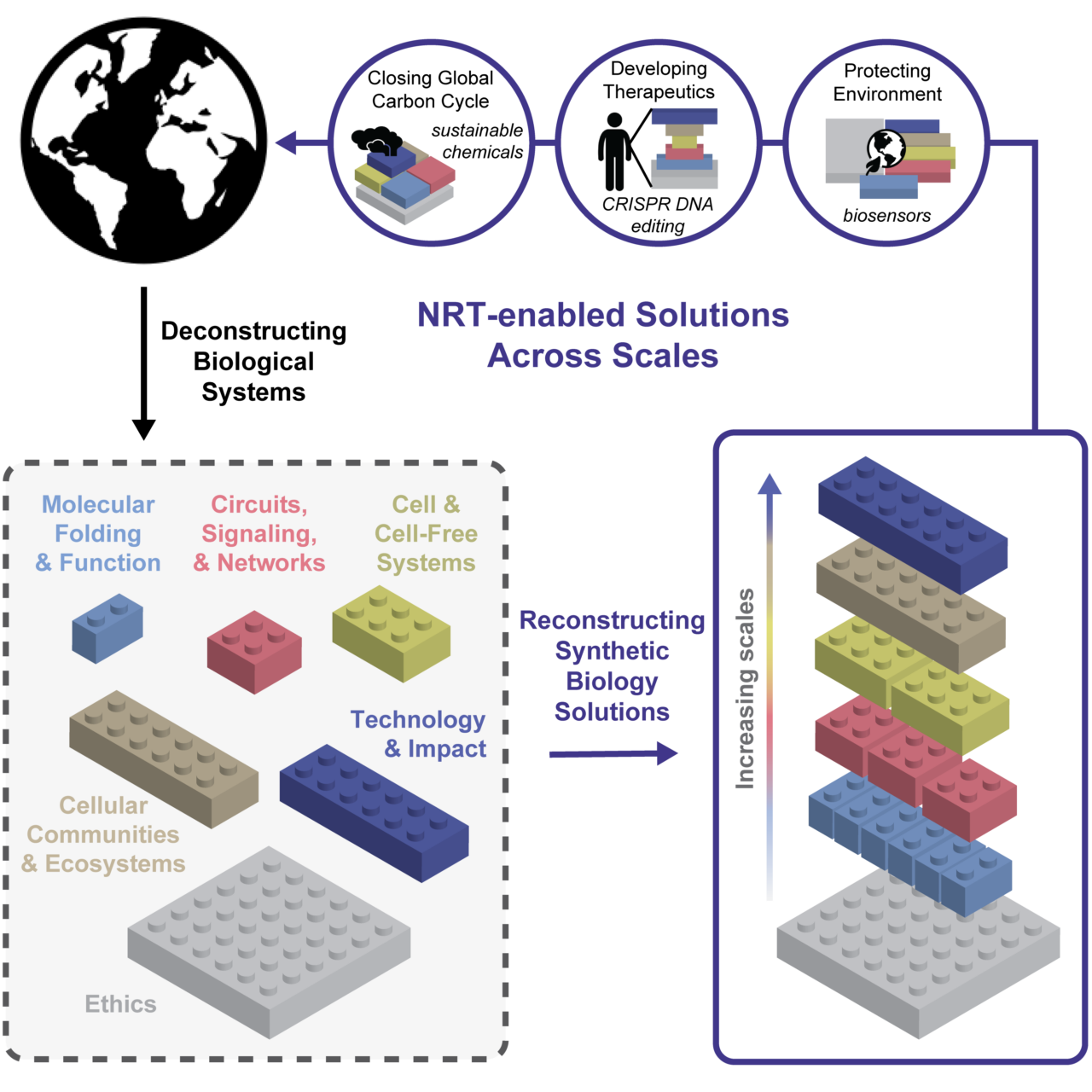
Understanding and harnessing the rules of life across multiple scales.
Life presents an enormous diversity of biological function that spans multiple spatiotemporal scales. These functions—-from the abilities of cells to synthesize small molecules, remediate environmental contaminants, build and maintain ecosystems, and differentiate to protect our immune systems—-have great potential to become components of sustainable solutions for meeting pressing global challenges. For example, synthetic biology research has led to engineered biological systems that can synthesize fuels, pharmaceuticals, and foods from sustainable feedstocks, act as smart therapeutics to cure diseases, and help balance the global carbon cycle. Even the seemingly simple example of cellular synthesis of products from sustainable feedstocks requires understanding and synthesis of diverse phenomena, including the underlying reaction chemical kinetics (chemistry), enzyme biophysics and substrate transport (physics), genetic regulation of enzymes and cellular physiology (biology), reactor vessel scale-up (engineering), and technoeconomic analyses (business).
Learn more about the scales frameworkProgram Components
SynBAS consists of several key components and activities
Our training program consists of an integrated curriculum, interactive workshops, and an innovative experiential learning project. Trainees begin with coursework that teaches students how to deconstruct synthetic biology technologies along the different scales, identifying the design principles required at each scale and the challenges between scales that must be overcome, for that application. Trainees then choose elective courses that emphasize details at particular scales related to their research. Through a suite of workshops, cohort discussions, experiential projects, ethics conversations, community activities, and more, trainees will learn what it takes to be synthetic biologists.
The SynBAS program consists of 5 components:
- Courses in Synthetic Biology. The synthetic biology core curriculum consists of a required case-study course on deconstructing biological function across scales. Elective courses along two different scales and chosen by the students provide rigorous training in the fundamentals of physics, chemistry, biology and ethics need to understand biological function at a particular scale. Coursework is typically started before applying to the program and completed while a trainee.
- Research Design and Communications Workshops. These monthly workshops provide a range of training opportunities to student. The research design portion of the workshops teach students how to frame research questions, develop specific hypotheses, design experiments to test those hypotheses, and analyze data to provide informative conclusions. The communications portion of these workshops give formal instruction on science communication, training on data visualization and presentation, and practice giving presentations. These workshops are also used to create community forums on the ethics of synthetic biology research. Students participate in this workshop during the funded period of their traineeship.
- Python Programming Workshop. This workshop introduces students to programming fundamentals with Python programming, and with the use of scientific tools for data analysis, simulation and data visualization. Students typically take this workshop the summer before their traineeship starts.
- Experiential Learning. In their second year of support, NRT trainees will gain hands-on experience leveraging their training to impact broader society. Trainees will choose a topic area underrepresented in traditional graduate training such as Education, Entrepreneurship, or Policy. Trainees will then develop and execute a custom, open-ended experiential project in their chosen area. All experiential projects will be presented at the annual CSB retreat to communicate results to the broader community. Several examples are included here, but trainees are welcome to explore several options. Trainees could:
- Devise and conduct a summer outreach module for undergraduate and/or high school curricula in synthetic biology with embedded high-school teachers.
- Work with Northwestern Kellogg School of Management faculty and students to develop and pitch a synthetic biology-based business model to a panel of local entrepreneurs, which would give trainees the skills to translate discoveries into society.
- Critically analyze and discuss policy considerations surrounding synthetic biology and develop a policy product such as policy brief or public workshop on bioethics.
- Co-Mentored Thesis Research. Trainees are expected to find a co-mentor for their thesis research during the first year of their support. An ideal co-mentor is one that can provide important expertise in a scale that is complementary to that of the trainee’s primary advisor.
- Community Activities. The CSB conducts a number of community activities including an annual retreat and regular get togethers. Trainees play an active role in shaping and implementing these programs.
The Application Cycle
How to Apply
Application Deadlines
The deadline for the receipt of applications: July 1, 2024
Decisions expected: Mid-August
Fellowship appointment will begin in September, at the start of fall quarter.
Eligibility
- All current graduate students can apply.
- There are no requirements on the year of graduate study for applicants if the program requirements can be met.
- Funded and unfunded training program slots are available for participation. Due to NSF requirements, only US citizens or permanent residents can receive funding support.
Requirements
In your application, you will be asked to include the following:
- Applicant Statement
- Motivation for participating in this NRT.
- A description of your PhD research plans.
- A personal biography.
- Your plans for fulfilling the program requirements.
- Agreement to Trainee Expectations Document
- Letter of Support from Your Advisor



2024-2025 Cohort

Katy Bond
PhD Student

Juliana Feng
MD/PhD Student

Ethan Halingstad
PhD Student


Natalia Mendonca
PhD Student

Christina Thaggard
PhD Student

Amy Weiss
PhD Student
2023-2024 Cohort
2022-2023 Cohort

Delfin Buyco
PhD Student

Emanuelle Grody
PhD Student

Elizabeth Johnson
PhD Student

Tyler Lucci
PhD Student
SynBAS Trainees
2021-2022 Cohort

Gauri Bora
PhD Student

Dylan Brown
PhD Student

Vivian Hu
PhD Student

Brett J. Palmero
PhD Student

Claire Phoumyvong
PhD Student










